You will be redirected to:
Context
Clinical trials produce a massive amount of data. In most cases, the challenge lies less in the volume of data to process, but more in its diversity. The sources of data are indeed numerous and do not stop increasing. We first think about the data collected via forms that the physicians have to fill in, after each patient visit to a site, via an electronic data capture system (EDC), or the report forms produced by laboratories. But nowadays, clinical trials are getting more patient centric, and electronic patient reported outcomes (ePRO) are collected via web forms or mobile phones. Furthermore, the amount of image data (scanner, MRI, biopsy...) is also growing; and the development of non-invasive wearables collecting signals (heart rate, blood oxygen level...) is also increasing the need for proper data storage and manipulation systems.
Besides, with the diversity of data comes along more metadata, i.e. the data about the data, which needs to be properly stored and managed. Even if many organizations are pushing for standardization, it is far from being the norm at the moment, and data managers often have to pool and map data to a common format to make analysis possible.
These changes put more and more pressure on the shoulder of the clinical data managers, the goldsmith of clinical data, who have to identify, and configure the relevant technologies to make the clinical data ready for submission to authorities and available in the right format for bio-statisticians, physicians and other investigators. In addition, all the clinical data generated during a clinical trial, the Trial Master File (TMF), must be properly stored, organized and made available. This must be done for regulatory reasons, but also to enable cross-study analysis or optimize clinical trial operations.
Technological trends
More integrated services into clinical trial platforms
Clinical data managers spend a lot of their time transferring data from one application to another. These data transfers are increasing nowadays, since a more and more diverse portfolio of services is required to properly conduct a clinical trial. However, those data transfers are often manual, and thereby time consuming and prone to errors. It is indeed not unusual that a clinical data manager, to perform some data transformations: needs to connect to an EDC system deployed on cloud A; get some study data out by querying a database or exporting flat files; and then needs to connect to another server on cloud B to use SAS or R packages parameterized by Excel files. This is too complex! Data transformation capabilities should be embedded into the EDC. Furthermore, manual data transfers prevent any near real time data treatments which are definitely in need, e.g. for pharmacovigilance monitoring.
To automate/eliminate those manual data transfers, many vendors combine different applications into an integrated platform, where different services are interconnected and can operate seamlessly. The number and maturity level of services integrated vary from one vendor to another, but these platforms are currently under strong development. Some vendors provide a full clinical trial service coverage: integrating a clinical trial management system (CTMS), an EDC system, an eTMF, and advanced reporting capabilities; whereas others started with a simple EDC system and extend it with CDISC-SDTM mapping capabilities or pharmacovigilance reporting systems. In addition, with the rise of data coming from labs and wearable devices, specific data ingestion and transformation tools are also integrated.
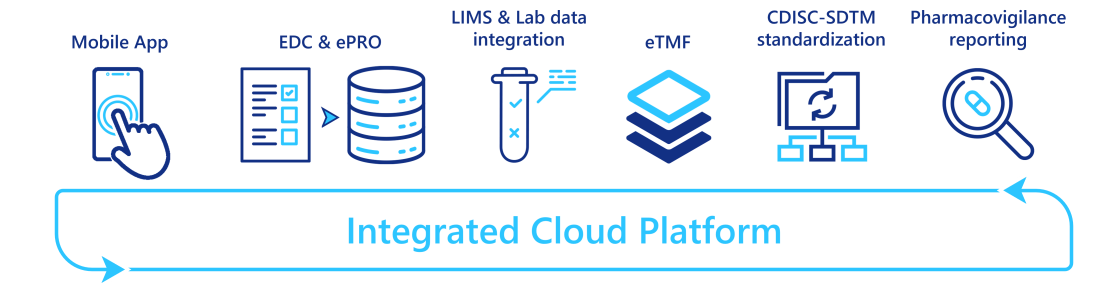
By providing a single access point to a myriad of services, these integrated platforms reduce the tedious and manual data transfers between applications, and so free time for the clinical data managers which can then be re-invested in more valuable tasks.
Clinical metadata repositories (CMDR)
Metadata is of crucial importance when one wants to interoperate technologies and promote reuse of codes. It is particularly important for data managers who need to design the data transformation flow, from the data capture by the EDC system, to the CDISC SDTM data format required for submission to authorities. This flow consists of two main tasks: configuring the EDC system based on the study protocol; and creating the relevant data outputs with the right format (CDISC-SDTM, CDISC- ADaM for statistical analysis, reporting tool formats). The latter, often referred to as "data mapping", is particularly tedious and time consuming. It often consists in writing codes in SAS or other data manipulation technologies.
One solution is to promote standardization which allows for reuse of configuration parameters and codes, and therefore reduces the workload in testing newly implemented codes. However, reusing the configuration of previous clinical trials may not be as easy as it sounds. Indeed, copy-pasting from previous trial designs is a process prone to errors when metadata are scattered across several parameter files. Actually, a data manager may not even know where to find the relevant past trial information.
A clinical metadata repository (CMDR) is a software platform capable of storing and updating/modifying clinical trial metadata:
configuring settings from EDC systems such as information about the visits, forms and questions (CDISC-ODM standard);
the different versions of CDISC-SDTM standard or your own standards;
data mapping configuration.
It also enables automatic import and export of metadata from compatible EDC systems (see figure below).

By gathering all these standards and providing access to past studies metadata, CMDRs facilitate the re-use of past study configurations. Indeed, a clinical data manager can easily search for similar study metadata, make a copy of them, modify and complete them; and ask for approval without leaving the CMDR platform. Some CMDR platforms even enable the visualization of the forms/questions, so the study design can be performed without opening the EDC system. Besides, since past metadata studies have already been validated, CMDR platforms also mitigate the testing workload.
Moreover, by providing a single point of truth across your organization, CMDRs enable seamless collaboration across teams and countries. Like most software platforms in this domain, they also provide traceability control (who made updates to what and when), access control, and document version control.
Leverage NLP techniques to promote automation
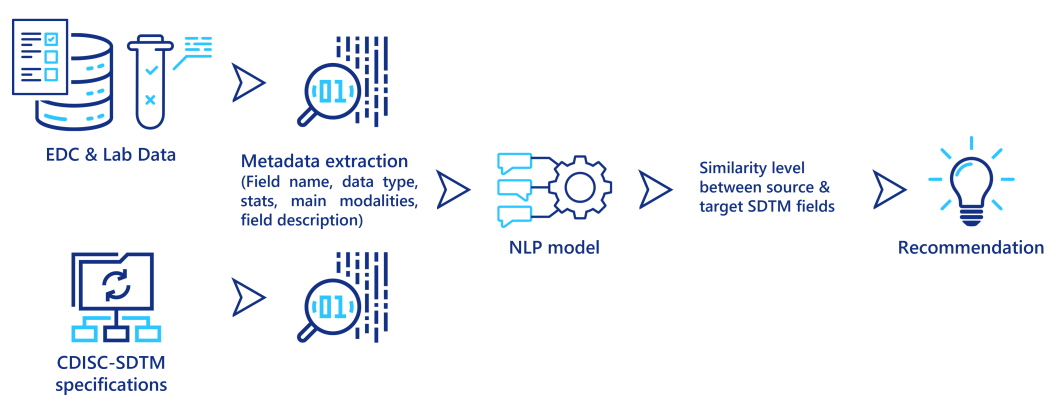
Natural Language Processing (NLP) techniques are becoming more and more powerful at extracting relevant information from textual data. In particular, these techniques are capable of predicting a similarity measure between two different textual descriptions, to determine if they both describe the same concept, object or entity. They can be used to assist clinical data managers when they map the data from its source system (EDC output, lab data) to CDISC-SDTM. Some metadata is first extracted from the source data and from the SDTM specifications, then for each field in the source data, an NLP model computes a similarity score to the fields in the SDTM standards, and recommends the similar ones to the data managers (see figure below). Even if this approach still requires a final validation from a clinical data manager, it could significantly reduce the time required to perform these tedious mapping tasks.
A second application of NLP techniques is document classification. Here the document type is automatically inferred from its content. This is useful to classify the myriad of documents generated during a clinical trial into their corresponding folder in the TMF, and thereby ensuring its quality. It can be particularly interesting to ingest past clinical trial data stored in paper format after the scanning and optical character recognition (OCR) steps.
A third technique in NLP is Named-Entity Recognition (NER). The idea here is to recognize/locate named entities (word or group of words) and classify them into categories (name, address, patient personal information, drug, posology, units, eligibility criteria...). NER techniques embed domain specific taxonomies or ontologies and could be used to automatically extract information from study protocols. This technology as well as the promotion of protocol templates/standardization can pave the way to automatic EDC configuration from the study protocol.
Those NLP techniques mainly rely on machine learning models specifically designed for text processing. Like any model, its performance depends on the quality and amount of data used to train it. To begin with, you may not have enough data to train a good model adapted to your context, but a model properly integrated into a software platform could be re-trained each time a user corrects its prediction, and so the model improves its predicting power on the way.
Even though the actual benefit of such models should be carefully assessed, they are very promising to support and automate tasks in clinical data management.
How to take advantage of these technological advances?
To properly embrace those technological trends, it is crucial to analyze the current processes and the technological stack in place in your organization, in parallel to your current and future needs. Then, you can determine what is the best way forward; from identifying the appropriate vendors to complement your existing technological stack, to develop some proof of concepts to assess the feasibility and the potential impacts of these technologies.
Leveraging our experience and capabilities as a CRO and as a digital transformation company, Keyrus can help you identify your needs and constraints and help you shape an action plan during a scoping study. With a diverse workforce of clinical data managers, data architects, data scientists and clinical project managers, we can fully support you in your technological transition.
